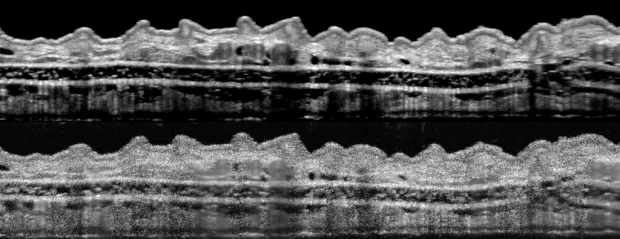
Photo courtesy de la Zerda lab: The upper image, showing a section of a mouse’s outer ear, is considerably en-
hanced by the de la Zerda group’s refinements compared with one obtained by conventional technology (lower image).
Stanford Medicine News Center - June 20th, 2017 - by Bruce Goldman
You may not have heard of optical coherence tomography, or OCT. But if you’ve visited an ophthalmologist recently, chances are your eye came within an inch or two of a scanning device employing the technology. Tens of thousands of these devices are in place in doctors’ offices, where they’re widely used to check for eye diseases.
Now, Stanford University scientists have figured out how to retrofit these high-performance machines with off-the-shelf components, increasing OCT’s resolution by several-fold and promising earlier detection of retinal and corneal damage, incipient tumors and more.

Photo by Norbert von der Groeben: Adam de la Zerda and
his team have devised a way to improve the quality of
images obtained through optical coherence tomography.
The relatively simple, low-cost fix — entailing a pair of lenses, a piece of ground glass and some software tweaks — erases blemishes that have bedeviled images obtained via OCT since its invention in 1991. This improvement, combined with the technology’s ability to optically penetrate up to 2 millimeters into tissue, could enable physicians to perform “virtual biopsies,” visualizing tissue in three dimensions at microscope-quality resolution without excising any tissue from patients.
In a study published online June 20 in Nature Communications, the researchers tested the enhancement in two different commercially available OCT devices. They were able to view cell-scale features in intact tissues, including in a living mouse’s ear and a human fingertip, said the study’s senior author, Adam de la Zerda, PhD, assistant professor of structural biology. The study’s lead author is electrical-engineering graduate student Orly Liba.
Boosting resolution with minimal tinkering
“We showed that you can take effectively any OCT system out there and, with minimal changes, boost its resolution to the point where it can detect anatomical features smaller than the size of a typical cell,” de la Zerda said.
OCT is a billion-dollar business. Every year, more than 10 million OCT scans are performed to diagnose or monitor conditions from age-related macular degeneration to melanoma. The technology has been adapted for endoscopic use in pulmonary, gastrointestinal and cardiovascular medicine.
Somewhat analogous to ultrasound, OCT penetrates tissues optically instead of with sound waves. The device aims beams of laser light at an object — say, a tissue sample, or a patient’s eye — and records what comes back when light bounces off reflective elements within the sample or eyeball. Adjusting the depth of penetration, a user can scan layer upon layer of a tissue and, piling virtual slices of tissue atop one another, assemble them to generate a volumetric image.
But to this day, OCT continues to be plagued by a form of noise that, unlike the random noise generated by any sensing system, can’t be “washed away” simply by repeatedly imaging the object of interest and averaging the results with a computer program.
The noise generated by OCT, called “speckle,” is an inherent feature of the architecture of the object being viewed and the unique properties of laser light.
A photon isn’t a mere particle. It’s also a wave whose power waxes and wanes as it travels, similar to an ocean wave heading toward the shore. When two waves collide, their combined height at the moment of their collision depends on whether each was at its peak, its trough or somewhere in between.
When photons get out of phase
The photons comprising a beam of laser light are in phase: They share the same wavelength, with their peaks and troughs occurring in sync. But when these photons bounce off of two separate surfaces — say, two closely situated components of a cell — the length of their return routes differs slightly, so they’re no longer in phase. Now, they can interfere with one another just like intersecting ocean waves. They may cancel each other out, creating a false-black speckle on the resulting image. Or they may reinforce one another, creating a false-white speckle. If the speckle-generating components’ positions are fixed, as is the case in most tissues (circulating blood being one exception), those same speckles will pop up in the same places on every successive OCT scan.
“Other researchers have tried various fixes, such as scanning repeatedly at different angles or from consecutive adjacent positions or with shifting wavelengths, or ‘removing’ the speckles using computer post-processing,” de la Zerda said. “But the result is always the same: a blurred image.” It’s like covering up freckles with a coat of makeup: a smoother appearance, at the cost of lost detail.
In principle, if you could reach in with a molecular tweezers and move one of those two interfering components just a tiny bit, you would change the speckle pattern. But you can’t. However, the Stanford scientists found a way to do essentially the same thing, optically speaking.
“We wanted to make the speckles dance, so they’d be in a slightly different pattern each time we scanned the tissue,” Liba said. “And we found a way to do it.”
Creating a virtual image
By positioning a couple of additional lenses in the OCT device’s line of sight, the investigators were able to create a second image — a holograph-like exact lookalike of the viewed sample that appeared elsewhere along the line of sight, between the added lenses and the sample. By inserting what they call a “diffuser” — a plate of glass they’d had roughened by randomly etching tiny grooves into it — at just the right point in the line of sight and methodically moving it between each round of repeated scans, they achieved the optical equivalent of shifting the geographical relationship of the sample’s components just a tiny bit each time they scanned it.
Now, averaging the successive images removed the speckles. The Stanford team used the resulting enhanced capability to acquire detailed, essentially noise-free images of a living, anesthetized mouse’s ear.
“We saw sebaceous glands, hair follicles, blood vessels, lymph vessels and more,” Liba said.
They also obtained high-resolution images of a mouse retina and cornea. And an incision-free look at the fingertip of one of the study’s co-authors let them see an anatomical feature never before glimpsed with OCT: Meissner’s corpuscle, a nerve bundle responsible for tactile sensations.
The technological advance gets around a 25-year-old problem that has persistently limited OCT’s diagnostic capabilities, de la Zerda said.
The work is an example of Stanford Medicine’s focus on precision health, the goal of which is to anticipate and prevent disease in the healthy and precisely diagnose and treat disease in the ill.
De la Zerda is a member of the Stanford Cancer Institute and of Stanford Bio-X and is a Chan-Zuckerberg Biohub investigator. Liba is a Bio-X fellow.
Stanford’s Office of Technology Licensing has applied for patents on intellectual property associated with the findings in the study.
Other Stanford co-authors of the study are former postdoctoral scholars Matthew Lew, PhD, and Debasish Sen, PhD; graduate student Elliott SoRelle; research assistant Rebecca Dutta; professor of ophthalmology Darius Moshfeghi, MD; and professor of physics and of molecular and cellular physiology Steven Chu, PhD.
The work was funded by the National Institutes of Health (grant DP0D012179), the National Science Foundation, the U.S. Air Force, the Claire Giannini Fund, the Damon Runyon Cancer Research Foundation, the Susan G. Komen Foundation, the Mary Kay Foundation, the Donald E. and Delia B. Baxter Foundation, the Skippy Frank Foundation, the Center for Cancer Nanotechnology Excellence and Translation, and Stanford Bio-X.
Stanford’s Department of Structural Biology also supported the work.




